The Workflow
The basic workflow of GLAD involves the following steps
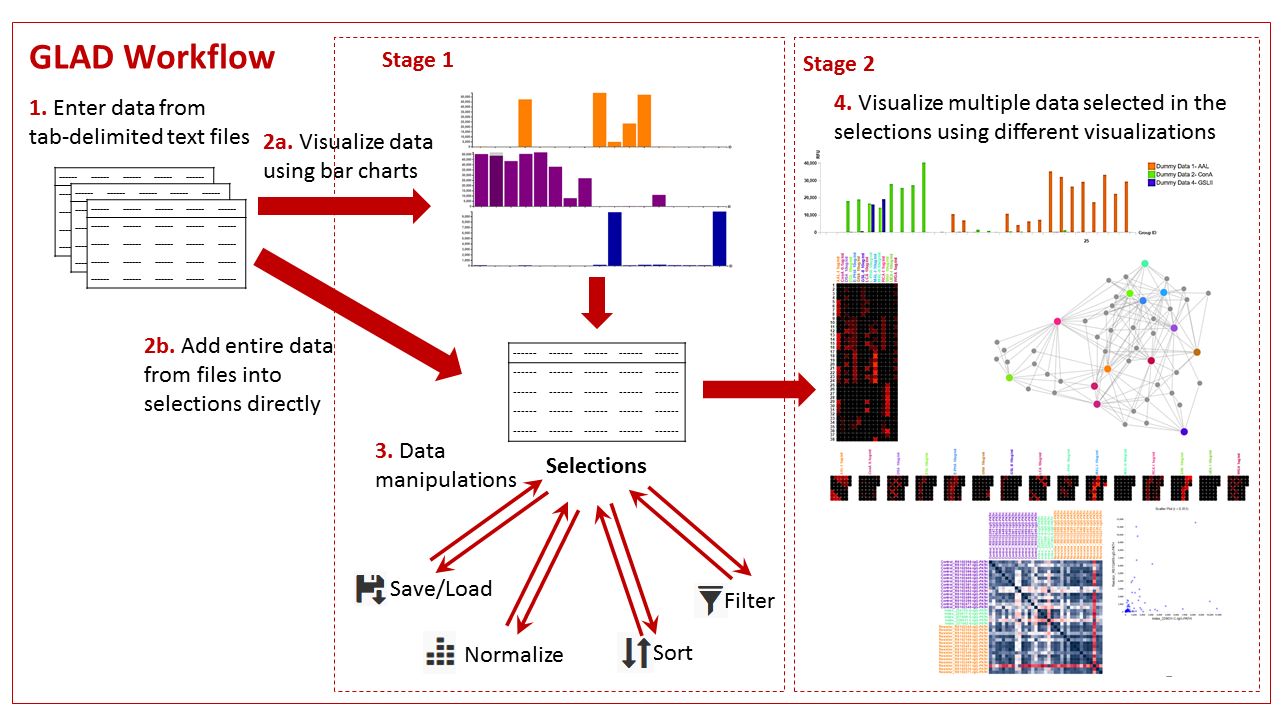
- Entering the data into the system in the correct format.
- Select the data into the program in one of two ways:
2a. Visualize the data in the form of bar charts and select the data which you would like to save together in “Selections”.
2b. Alternatively, the entire data file can be added to the “Selections” directly. - Manipulate the aggregated “Selections” data by either:
- Saving or loading it
- Normalize the data
- Sorting the data in particular order
- Filtering the data
- Present the data using one of the Stage 2 visualizations such as grouped bar charts, heatmaps, calendar heatmaps, force graphs, and correlation maps
The glycan structure information can also be viewed at any time by simply moving the pointer over the data points such as the bars in bar charts and the tiles in heatmaps.
The Interface

Above is an outline of the GLAD interface. It has 7 major regions which the user should be aware of.
- Users can load data which we make public, or their own data provided the file is in the correct format for GLAD
- Users can change the color representation of the data which they are loading. More details on colors can be found here <insert link>
- Users can display data as bar charts which can be used to select specific data for analysis in stage 2. Multiple bar charts can be produced so as to compare data and make selections.
- Users can save / load selections, filter, sort or normalize the selections.
- Users can use the selected data from multiple array experiments to visualize the data as one of the various stage 2 visualizations offered such as:
- Grouped Bar Charts
- Heatmap
- Calendar Heatmap
- Force Graphs
- Correlation Maps.
- Clicking on the Glycan Structures button opens up a drawer which can show you the glycan structure based on the CFG name.
- This is where all the data visualization happens.
Additional settings for different items are denoted by the settings cog-wheel ![]()