One of the strongest advantages GLAD offers in order to improve glycan microarray data visualization, is the ability to filter the data in an instantaneous way.
Filtering Data
GLAD allows you to filter data by:
Filter by Query
To filter selected data:
- Make sure you have loaded data as selections.
- Click on the
 button.
button. 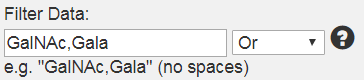 In the input field type in the filtering query . The input query would be based on text which is present in the Name, Name2, ID or Data Titles which were originally entered in the data files. For example:
In the input field type in the filtering query . The input query would be based on text which is present in the Name, Name2, ID or Data Titles which were originally entered in the data files. For example:
- To filter by Glycan Name: to filter data for all glycans containing sialic acid, you could enter “Neu5Ac” in the input field.
To perform a terminal monosaccharide filter include ‘T:’ before the monosaccharide, for example, the query ‘T:Fuca,T:GlcNAc’ with the option And will filter all glycans which have terminal Fuca and terminal GlcNAc. - To filter by IDs: type in the id numbers separated by commas. You can even type in range of glycans. For example, “10-15,30-40” will filter out data for glycans with IDs 10-15 or 30-40.
- To filter by Sample Name/Data Titles: to filter the data for a specific lectin in a selections file with multiple lectins for example AAL you would put in “AAL” in the input field.Note the query string does not need to contain full names of the samples or glycans. It just needs enough information to differentiate between them.
- To filter by Glycan Name: to filter data for all glycans containing sialic acid, you could enter “Neu5Ac” in the input field.
- You can enter multiple filtering criteria separated by commas in the input field and choose one of the filtering options such as Or, And, Not Or and Not And from the drop down next to it. For example if you would like to see the data for Glycans containing both GalNAc and Galα, your query would be “GalNAc,Gala” and your filtering option would be of the type Or. This means that all data for glycans which contain either GalNAc or Gala would be filtered out.
More details about the filtering options is given below. - Choose how you would like to filter the data:
- Filter Glycans by Name
- Filter Glycans by Name2 (alternate name filtering)
- Filter Glycans by ID numbers
- Filter Samples (filters by Data Titles)

- The original selections (i.e. before filtering) can be restored after filtering by clicking on the Restore Selections button
 which appears next to the Filter Data button.
which appears next to the Filter Data button.
Understanding the Different Filtering Options
GLAD uses mathematical set and logic operations of unions, intersections and complements to give you Or, And, Not Or and Not And options.
Assume that we are filtering by Glycan Name and the glycan name is made up of A, B, C and D.
The diagram shows what the Or, And, Not Or and Not And options would filter out shaded in blue.

In this diagram:
- A Or B : would filter out data which contain either A or B in the field being filtered. Hence it is equivalent of A∪B in set theory.
For example: A, AC, BAC, CBA, CAD, DAC…. (either A or B must be present)
- A And B : would filter out data which contains both A and B in the field being filtered. Hence it is equivalent of A∩B in set theory.
For example: CBA, CAB, BAD, DBA, DAB … (both A and B must be present) - Not A Or B : would filter out data which contains neither A nor B in the field being filtered. Hence it is equivalent of (A∪B)C in set theory.
For example: CD, DC, C, DCC, CDD, CDC … (cannot have A or B)
- Not A And B : would filter out data which does not contain both A and B in the field being filtered. Hence it is equivalent to (A∩B)C in set theory.
For example: BDC, BD, CA, CDA, ACD …. (cannot have both A and B)
Filter by Cutoff and Filter by Rank
GLAD allows users to filter data by either Cutoff % of top binders or Top rank numbers. The following options need to be filled in:
- Top % Binders or Top Rank number should have the threshold percentage or number respectively.
- Keep data for all samples if above cutoff for even single sample: This option allows users to keep data for the glycan from all samples even if the glycan shows binding above the threshold for a single sample. This allows the user to compare the data for the glycan among all samples even if it binds to one.
- Perform sort by ID after filtering based on cutoff: This option sorts the data by ID after filtering so as to restore the order. The the filter function would go sample by sample and reorder the glycans by the sample.
Additionally, the Filter by Rank has an option to chose the Rank Order either:
- Descending i.e. lower rank number = stronger binder. For example, rank 1 is strongest binder.
- Ascending i.e. higher rank number = strong binder. For example rank 100 is stronger binder than rank 80.
This depends on how the rank was calculated in excel or other software before.